
The growth of healthcare data
Globally, we’re seeing what is often talked about as an ‘explosion’ of healthcare data. In part driven by our new ability to generate digital data through technology like devices and wearables, and in part driven by a desire to improve healthcare provision through use of data, the data being generated and used is growing rapidly. According to one source, the healthcare industry generates around 30% of the world’s data, and there’s no sign of slowing down. It also predicts the compound annual growth rate of healthcare data will reach 36% by 2025 (significantly faster growth than manufacturing, financial services, or media and entertainment).
The healthcare data used in clinical trials is no exception to this trend. Recent studies have shown that a typical phase three oncology trial will generate an average of 3.6 million data points. This is three times the data collected by late-stage trials 10 years ago.
The potential value held in all the healthcare data now being routinely collected is widely recognised. It can:
- Be used to inform and improve individual patient care, for instance by flagging issues and recommended actions.
- Hold valuable insights on wider healthcare issues and patterns.
- Be used by healthcare systems to plan, improve and be more efficient with their services.
- Be used in the design and development of healthcare treatments and solutions (such as during clinical trials).
Healthcare data: Some challenges
But the ability to release the potential of healthcare data relies on a few key factors:
- Interoperability – The further use of healthcare data tends to rely on the transfer of data from one system to another.
- Quality – For data to be useful, it has to be accurate, reliable, up-to-date and complete (amongst other things).
- Consistency – The digitization of healthcare data is a relatively new phenomenon without a universally agreed approach. So, with the volume of data being collected across different devices, people, organisations and countries, there can be distinct differences in languages and terminologies being used.
More on each of these three factors below:
Interoperability
The Institute for Electrical and Electronics Engineering (IEEE) defines interoperability as “the ability of two or more systems or components to exchange information and to use the information that has been exchanged.”
The extent of interoperability is often described on four levels, including by The Healthcare Information and Management Systems Society (HIMSS):
- Foundational interoperability – Where one system or application can securely send and receive data from another. The receiving system does not necessarily need to be able to interpret the data.
- Structural interoperability – Where the format, syntax and organization of data exchange is defined at the data field level, allowing for interpretation of information.
- Semantic interoperability – Where systems can exchange, interpret and actively use information, for instance healthcare organisations with different Electronic Health Record (EHR) systems sharing data to improve and inform care.
- Organisational interoperability – Seamless communication of data within and between organizations, entities and individuals, with all the governance, policy, social, legal and organizational considerations in place.
The ability to achieve at least some level of interoperability has come a long way since the introduction of HL7 FHIR standards to many healthcare systems. This facilitates the packaging up of data in a standard format, and means that most EHRs now have a standard interface (or FHIR app) that allows for it to be connected to another system.
Quality
Achieving quality data collection has its own challenges and solutions, including because of a perceived lack of time and skills for quality data recording. But it’s also worth bearing in mind that, even when quality data entry is achieved, input does not necessarily equal output. Data quality can be damaged and information ‘lost in translation’, despite being able to connect systems at a basic level.
Consistency
Connected closely with quality, consistency in healthcare data is hard to achieve when there are countless possible ways of recording that someone has, for instance, breast cancer. Malignancy, carcinoma, sarcoma, CA – the variations go on. One way healthcare systems tackle this is by using standard reference ontology, terminology and coding systems. Yet there are hundreds of possible healthcare ‘languages’ like this to choose from. Different organisations use different languages, and within organisations, different languages are used for different purposes. A few of the most commonly used are:
- SNOMED CT (Systematized Nomenclature of Medicine — Clinical Terms) – This is viewed as the most comprehensive reference terminology in use internationally. It was mandated for use by everyone working in the NHS from April 2020. In 2013, the US government required that SNOMED CT be included in EHR systems in order for them to be certified for Stage 2 Meaningful Use (US regulations that incentivise interoperability in healthcare).
- LOINC (Logical Observation Identifiers Names and Codes) – A database and common terminology for identifying medical laboratory observations and clinical measures like vital signs. LOINC codes describe in detail how a particular lab test was done, containing six parts in a specific order: Component (the substance being measured); Property measured; Timing (the interval the test was taken); System (the origin of the specimen); Scale; and Method.
- ICD-10-CM – The US version of the International Classification of Diseases, created and maintained by the World Health Organization (WHO). Hospitals mainly use these codes for billing and reimbursement. Nationally and globally, the ICD is used to track morbidity and mortality statistics. Under ICD-10-CM, every disease or health condition is assigned a unique code three to seven characters long.
Data mapping: the solution
There are many facets to the three factors discussed, but you could say the solutions to interoperability, quality and consistency challenges meet in one place: data mapping.
“Data mapping involves ‘matching’ between a source and a target, such as between two databases that contain the same data elements but call them by different names. This matching enables software and systems to meaningfully exchange patient information, reimbursement claims, outcomes reporting and other data.” AHIMA’s HIM Body of Knowledge
Data mapping is key to creating a link between systems – one as the source of data, and the other the target – the system where the data needs to be. You can think of the process as creating a combined route map and bilingual dictionary, including all the data elements needed in the target system. The fields in the target system are matched to the relevant fields in the source system, and the terms and codes that may appear in the source are translated into the desired format for the target system.
Data mapping in clinical trials
As we’ve already mentioned, the quantity of data required for a typical clinical trial is growing at pace. Yet, currently, in a typical phase two oncology trial, around 70% of the study data already exists in the trial site’s hospital EHR.
A clinical trial’s sponsor (typically a pharmaceutical company) requires particular data to be complete in their clinical trial system (often termed Electronic Data Capture (EDC). The required data is detailed on Case Report Forms (CRFs). Some trials may only include a handful of CRFs. For others, each CRF could run to hundreds of pages, repeated for thousands of participants. Either way, hundreds of pages of data will be collected per patient during the course of a clinical trial and a patient’s multiple visits to clinic.
As well as data specific to the disease being treated, examples of some of the basic information detailed on a CRF include:
- Date of visit
- Blood pressure
- Pulse
- Lab results
- Respiration
- Height
- Medications
Mappings ensure that data taken from the source system (typically an EHR) are transferred to the target system (typically an EDC) in a way that it is accepted and recognised. For example, ‘medication start date’ could be recorded as ‘dd mm yyyy’ on the source system, but can only be accepted as ‘yyyy mm dd’ on the target system. Similarly, the medication’s route of administration could be recorded on the source system as full words, such as ‘oral’, ‘intravenous’, ‘inhalation’, whereas the target system accepts these entries using abbreviations, such as ‘PO’, ‘IV’ and ‘INH’. With thorough and accurate data mapping, the information can be translated to allow for a meaningful transfer of data from one system to another.
Challenges to data mapping in clinical trials
The process of getting there, however, is not always as straightforward as it may sound. Typically, many of the same forms are used repeatedly by a trial sponsor but mapping data accurately can be a relatively manual, intricate exercise that is open to errors.
Some of the challenges include:
- Hospitals may use a mixture of standard terminologies, as well as their own vocabularies.
- Sponsors may, similarly, require data to use a mixture of standard terminologies and their own vocabularies.
- Multiple hospitals may be taking part in a clinical trial – thus the data needed is held on multiple EHRs. In some cases, even within a single hospital there may be multiple EHRs in use across different departments.
- Trial protocols (and the questions on the CRFs) can be changed both leading up to, and mid-way through, a clinical trial.
- The ‘joining’ of the two systems can be facilitated by HL7 FHIR standards and apps, but there can be variations in the way hospitals configure their EHR and FHIR settings.
- Crucial data may be held in unstructured (free text) fields in the EHR, which presents additional challenges.
- Data requirements change over time – an obvious example would be in the way race and ethnicity are recorded – categories can merge or become more specific, as well as having quite different approaches and requirements in different countries.
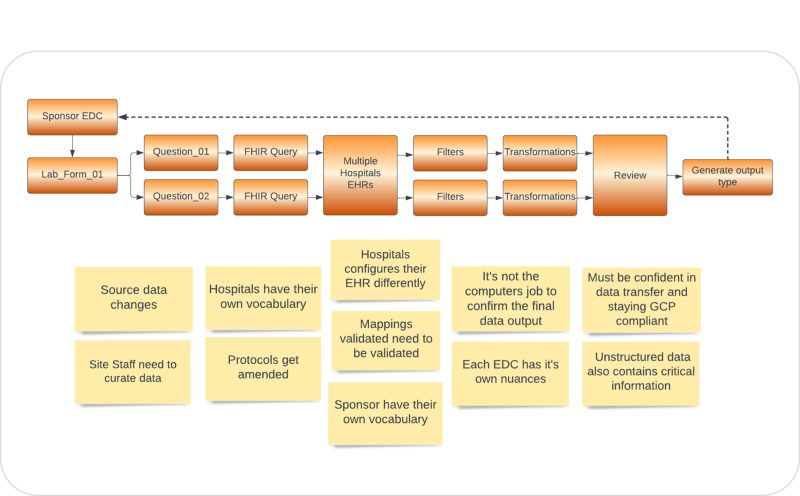
The ‘Great Mapping Challenge’
–
Archer – IgniteData’s healthcare data mapping genius
IgniteData is the company shaping the future of clinical trials. Through our innovative eClinical platform, Archer, we are enhancing interoperability between Electronic Health Records (EHR) and key research applications such as Electronic Data Capture (EDC).
We’ve been tackling the challenge of mapping healthcare data for years, and we understand the complexity of the task. Archer’s unique and sophisticated mapping engine has been purpose-built from the ground up – through deep consultation with both hospitals and sponsors – with this specific issue in mind. It streamlines and simplifies the task of mapping healthcare data from an EHR to a clinical trial system, eliminating errors and duplication of effort, and allowing clinical trials to go live faster.
Sponsors working with multiple trial sites can seamlessly merge data from all sites into their clinical trial system, mapping and re-mapping processes can be managed and tracked with ease, and previous mappings can be saved for re-use. Quality is ensured, while attaining huge time savings and efficiencies.
Find out more about how we make complex data mapping for clinical trials easy:
Follow Us
Share
More News
$8 million seed investment round turbocharges IgniteData’s growth
IgniteData secures £1/2 million Innovate UK grant for ground-breaking clinical trial data project
How EHR-to-EDC technology is sparking better collaboration in clinical trials
6 Common Data Challenges For Clinical Researchers
How to choose an EHR-to-EDC solution
Leading healthcare technology expert Steve Tolle joins IgniteData
Guest blog: Joseph Lengfellner on the challenges and solutions for clinical trials – Part 2
Guest blog: Joseph Lengfellner on the challenges and solutions for clinical trials – Part 1
How structured data is used in clinical trials
Evidence from Electronic Health Records-to-Electronic Data Capture live pilot study
IgniteData and Leading New York City Cancer Center Collaborate to Solve the Clinical Trial Data Transfer Challenge
Quality data and the EHR-to-EDC dream – where and why we need to focus our energies
UK clinical trials landscape: 4 big influencers
ZS’s Qin Ye on trends and driving innovation in life sciences

Meet Archer, the platform transforming EHR-to-EDC data automation
